Week 263
Foundation Models
In Week #263 of the Doctor Penguin newsletter, the following papers caught our attention:
1. Sleep Foundation Model. Polysomnography is a comprehensive overnight study that monitors brain activity, heart rate, breathing, and muscle activity during sleep. While typically used to diagnose sleep disorders, could these multimodal recordings predict a broader spectrum of future diseases?
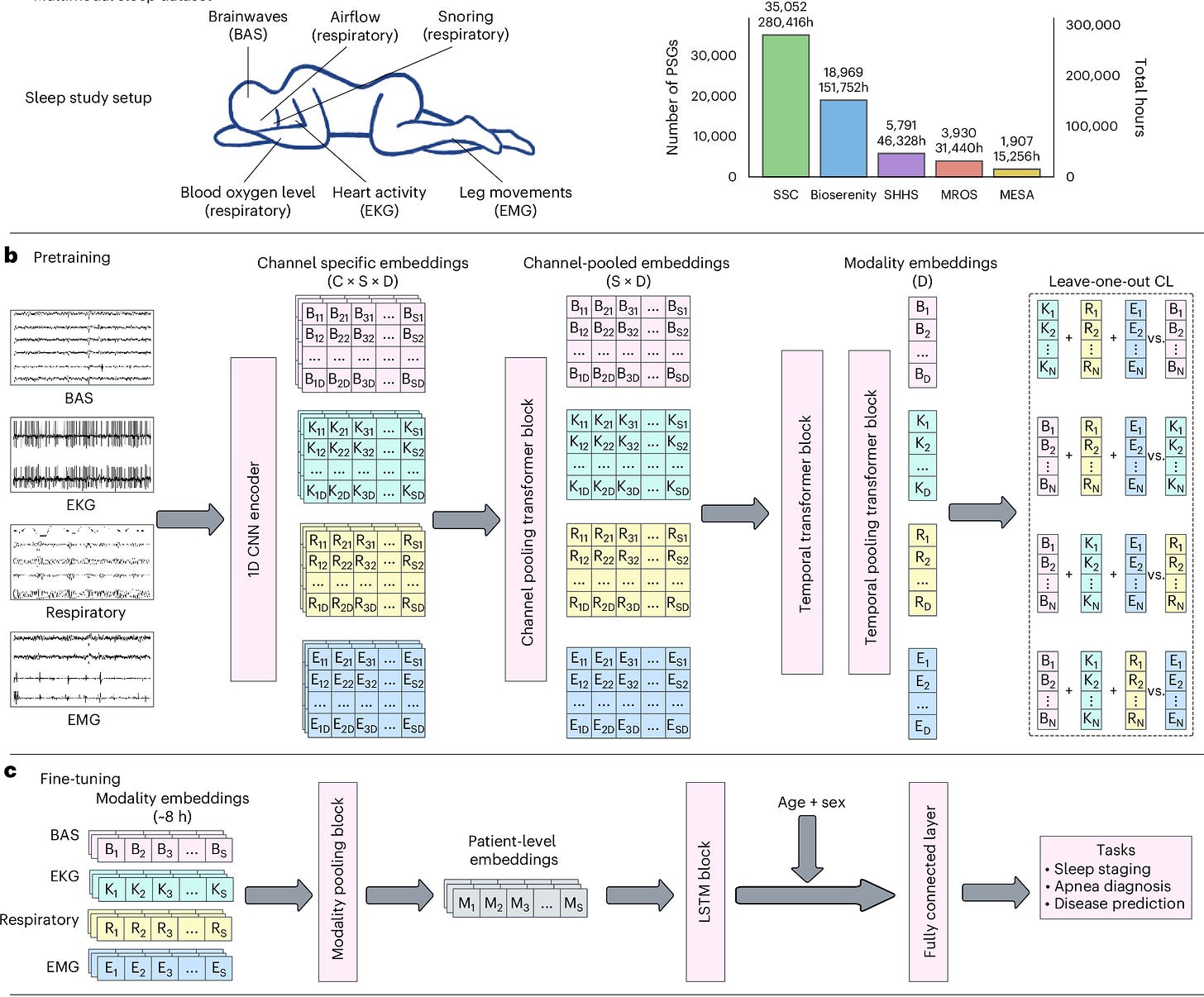
Thapa et al. developed SleepFM, a multimodal foundation model trained on over 585,000 hours of polysomnography data from 65,000 participants that predicts future disease risk from a single night of sleep. The model processes four signal modalities: brain activity (EEG/EOG), heart (ECG), muscle (EMG), and respiratory signals, by segmenting them into 5-second recordings (tokens) that are grouped into 5-minute context windows for processing by a temporal transformer. During pretraining, the model uses a novel leave-one-out contrastive learning approach, where each modality learns to predict the average embedding of the other three modalities. This strategy enables the model to handle missing modality during inference while learning shared physiological patterns across all signal types. During finetuning, a multi-task LSTM model was trained on the pretrained embeddings of full nights' recordings for disease prediction. SleepFM achieves C-Index of at least 0.75 for 130 conditions, including mortality (0.84), dementia (0.85), heart failure (0.80), and stroke (0.78). This study suggests that sleep's multimodal physiological signatures contain rich predictive information for early disease detection across cardiovascular, neurological, metabolic, and respiratory conditions.
Read paper | Nature Medicine
2. Echocardiography Foundation Model. Echocardiography is the most widely used cardiac imaging modality, using ultrasound to create real-time videos of the heart. Unlike natural videos, ultrasound videos are dominated by stochastic speckle patterns, depth-dependent intensity attenuation, and acoustic shadows. These artifacts vary across echocardiography acquisitions (patient body habitus, probe positioning, operator expertise, and equipment vendor) and bear no relationship to cardiac anatomy.
Munim et al. developed EchoJEPA, a foundation model for echocardiography that uses a latent predictive approach (predicting embeddings rather than reconstructing pixels) to learn robust cardiac representations from 18 million videos across 300,000 patients—the largest echocardiography pretraining dataset to date. While common pixel reconstruction methods for self-supervised video representation learning tend to memorize noise, latent prediction can ignore that and capture only temporally coherent anatomical structures like heart chambers and wall motion. This approach significantly outperforms pixel reconstruction methods, achieving 27% better performance on ejection fraction estimation in controlled comparisons, high sample efficiency (reaching 79% view classification accuracy with only 1% of labeled data versus 42% for the best baseline trained on 100%), and robustness to physics-informed acoustic perturbations simulating depth attenuation and shadows (degrading only 2.3% compared to 16.8% for competitors). Importantly, EchoJEPA's zero-shot performance on pediatric patients, a completely different population with smaller hearts and different anatomy, surpasses fully fine-tuned baseline models, demonstrating that latent prediction fundamentally captures more generalizable representations of cardiac anatomy rather than memorizing surface-level image statistics or acquisition-specific artifacts.
Read Paper | arXiv
3. Continuous Glucose Foundation Model. Continuous glucose monitoring (CGM) generates rich temporal data on glucose dynamics, but its full potential for predicting long-term health outcomes remains underutilized beyond immediate glycemic control.
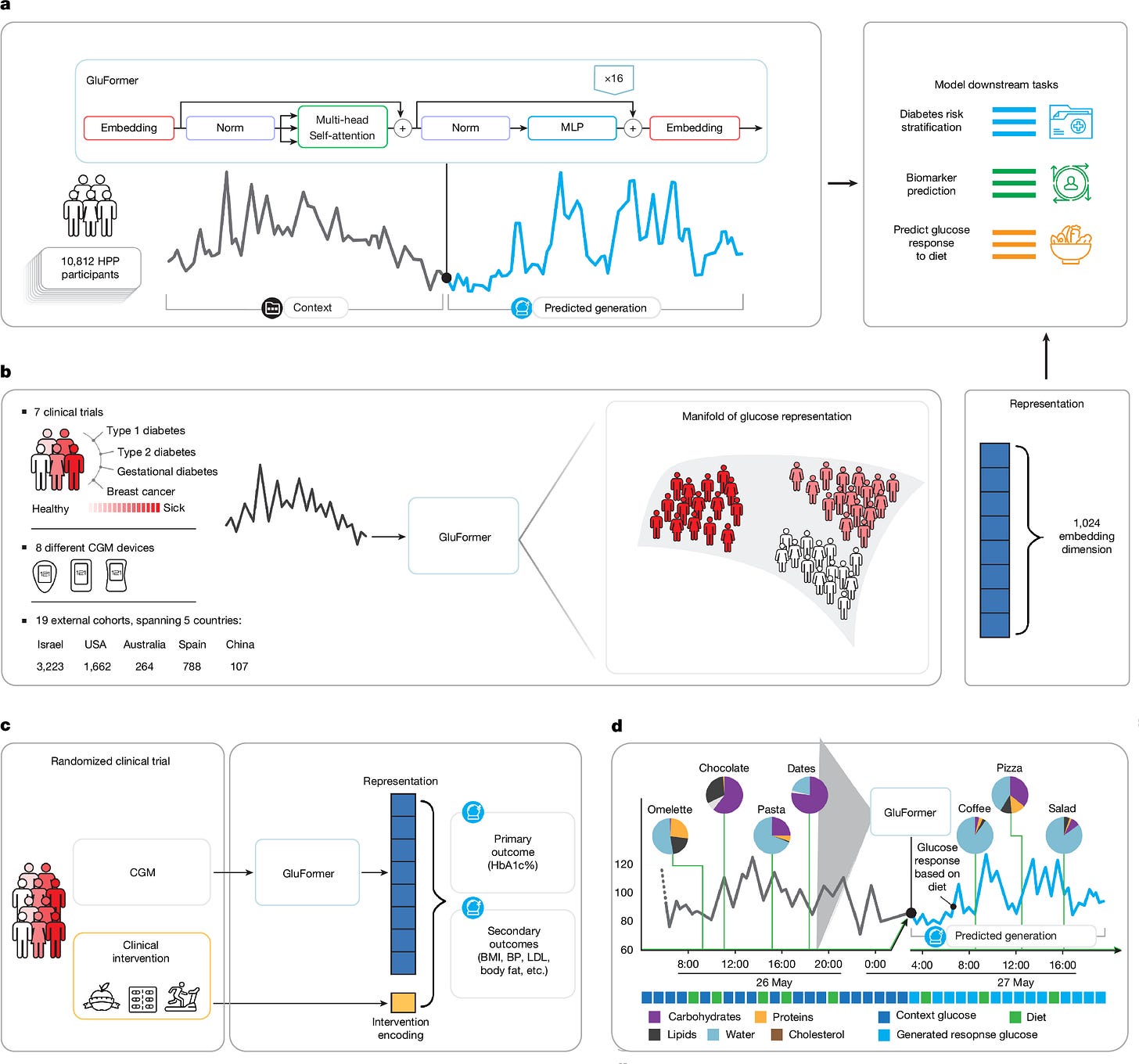
Lutsker et al. developed GluFormer, a CGM foundation model trained on over 10 million glucose measurements from 10,812 adults. The model uses a decoder-only transformer architecture where continuous glucose values (40-500 mg/dL) are discretized into 460 categorical tokens and trained autoregressively to predict the next token in the sequence. These learned representations generalized across 19 external cohorts spanning 5 countries, 8 CGM devices, and diverse conditions, including prediabetes, type 1 and 2 diabetes, gestational diabetes, and obesity. In a cohort with 11-year follow-up, GluFormer identified 66% of future diabetes cases and 69% of cardiovascular deaths in the top risk quartile, while baseline HbA1c stratification showed no significant association. This result suggests that the model could capture dynamic metabolic patterns missed by static biomarkers. A multimodal extension incorporating dietary data (calories, carbohydrates, proteins, etc.) aligned to CGM timestamps further improved predictions by using diet tokens as contextual input while training only to forecast glucose values. While improvements over standard metrics (GMI, iglu scores) were statistically significant but often modest for downstream prediction tasks, the model's consistent performance across populations and ability to stratify long-term risk from short-term CGM patterns suggest self-supervised learning on continuous physiological signals could complement traditional biomarkers for metabolic risk assessment.
Read Paper | Nature
4. Brain MRI Foundation Model. Brain MRI presents unique challenges for AI development due to its high-dimensional nature and heterogeneous acquisition protocols: a single brain MRI scan can produce different sequence types (T1-weighted, T2-weighted, T1CE, etc.) that vary across institutions and scanners, each providing distinct clinical information, with selection of sequences for analysis depending upon the clinical use cases. Despite the success of self-supervised learning in other medical imaging domains, there is a gap in developing MRI models that can generalize across different scanners, patient populations, and clinical tasks.
Tak et al. developed BrainIAC (Brain Imaging Adaptive Core), a foundation model for analyzing brain MRI scans across both healthy and diseased populations. Using self-supervised contrastive learning on 32,015 unlabeled brain MRIs, where the model learns to match different augmented views of the same brain region while distinguishing them from other regions, BrainIAC outperformed traditional supervised training and existing pretrained models across seven clinical tasks ranging from straightforward (MRI sequence classification) to challenging (predicting genomic mutations and patient survival). The model's advantages were most pronounced in limited data scenarios, maintaining strong performance with as few as 10% of available training data or even single examples per class, while also demonstrating robustness to common imaging artifacts. By learning generalizable representations from unlabeled data, BrainIAC enables AI development for rare diseases and difficult clinical scenarios previously infeasible due to limited labeled training data.
Read Paper | Nature Neuroscience
5. Non-contrast CT Foundation Model. Early detection of cancer through screening programs in asymptomatic populations significantly improved survival and outcomes compared to those diagnosed through standard clinical workflows. Non-contrast CT, particularly low-dose CT acquired during physical exams, offers a low cost and widely accessible imaging solution even in resource-limited settings.
Liang et al. developed OMAFound (carcinOMA Finder foundation), a 3D foundation model for simultaneous multi-cancer screening at both organ level and patient level using non-contrast CT. The model was trained and tested on 325,197 CT volumes from 151,386 patients across 10 datasets. Using paired CT mammography data, OMAFound demonstrated performance comparable to standard mammography-based approaches for breast cancer detection. The model also performed on par with established benchmark lung cancer screening models. In a prospective multicenter study of 21,601 patients, OMAFound showed balanced accuracy of 82.2% for breast cancer and 88.0% for lung cancer in females, and 86.1% for lung cancer in males. Additionally, when seven generalist radiologists were assisted by OMAFound, they showed significant improvements in sensitivity (mean increases of 38.9% for breast cancer, 16.0% for lung cancer, and 21.3% at the patient level) without compromising specificity.
Read Paper | Nature Health
-- Emma Chen, Pranav Rajpurkar & Eric Topol