Week 262
Continual Learning, Opportunistic Screening, Cell Segmentation, Protein Embedding Compression
In Week #262 of the Doctor Penguin newsletter, the following papers caught our attention:
1. Continual Learning. Medical AI models often perform well during development but struggle when deployed at new hospitals with different patient populations, equipment, and imaging protocols. Can continual learning help AI systems adapt across diverse clinical settings without sharing sensitive patient data?
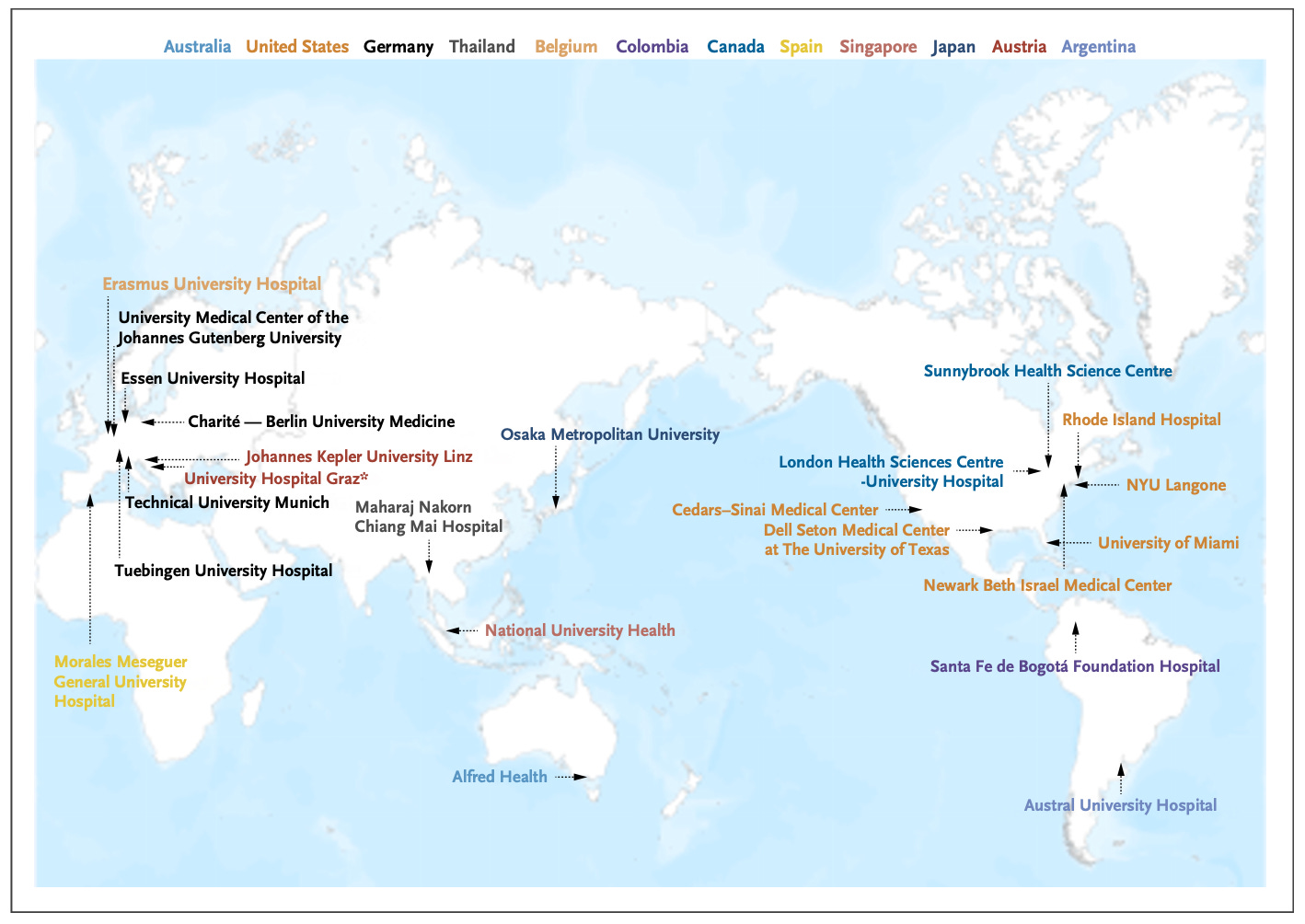
Chen et al. conducted an international retrospective study across 23 hospitals in 12 countries to test whether continual learning could improve an endotracheal tube (ETT) placement assessment model. Using 50 labeled chest radiographs per site, continual learning (where the model sequentially trains at each new hospital before deployment) helped the models achieve significantly lower ETT-carina measurement error (10.58 mm) compared to single-hospital fine-tuning (fine-tuning only at the deployment hospital) (12.49 mm) and the original model (16.39 mm). Performance improved progressively as more hospitals contributed to training. Notably, this approach requires no data sharing between institutions and is simpler than federated learning, making it practical for hospitals with limited IT infrastructure. The study demonstrates that medical AI models can accumulate knowledge across diverse international settings while preserving data privacy, offering a path toward more robust and generalizable clinical AI tools.
Read paper | NEJM AI
2. Opportunistic Screening. ECGs are not a conventional diagnostic tool for chronic liver disease (CLD), yet patients who undergo ECGs because of cardiac concerns may be at risk for CLD due to shared cardiovascular risk factors. Can AI enable earlier detection of advanced CLD in primary care using routine ECGs?
Simonetto et al. conducted a pragmatic cluster-randomized trial testing whether an ECG-based machine learning model can help detect advanced CLD in primary care settings. The trial randomized 98 primary care teams (245 clinicians, ~15,600 patients) to either usual care or receive notifications when the ML model flagged a patient as high-risk for advanced CLD. The primary endpoint was a new diagnosis of CLD with advanced fibrosis within 180 days of ECG. The intervention doubled the detection rate of advanced liver fibrosis (1.0% vs 0.5%, OR 2.09), with even stronger effects among ECG-ML-positive patients (4.4% vs 1.1%, OR 4.37). However, the overall diagnostic yield was lower than expected based on the 2-5% population prevalence of advanced CLD, which may be attributed to variable clinician adherence, with only 16% of patients with positive ECG-ML results receiving follow-up testing. Overall, the study shows that routine 12-lead ECGs could serve as an opportunistic screening tool for advanced CLD, since they’re already frequently obtained in populations at risk for liver disease due to shared cardiovascular risk factors like obesity and diabetes.
Read Paper | Nature Medicine
3. Cell Segmentation. Accurate cell segmentation is crucial for quantitative analysis and interpretation of various cellular imaging experiments. Although deep learning methods have made substantial progress, most models are specialists that work well for specific domains but cannot generalize across different imaging modalities or cell types.
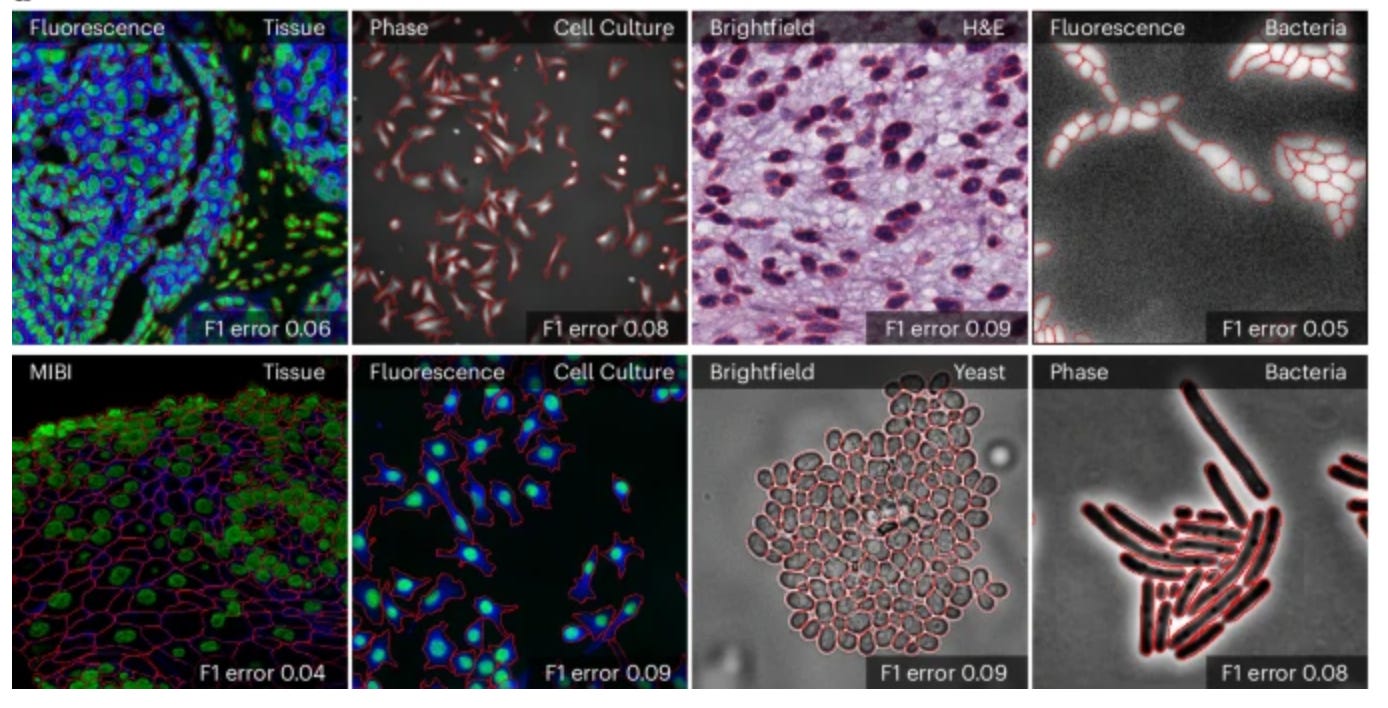
The "Segment Anything Model (SAM)" is a general segmentation model from Meta that identifies and segments any object in an image or video when given a simple prompt like a click, bounding box, or text. Building on top of SAM, Marks et al. developed CellSAM for universal cell segmentation across diverse cellular imaging data. To achieve this, they trained a vision transformer-based object detector that automatically detects cells and generates bounding boxes as prompts for SAM. CellSAM was trained on a comprehensive dataset spanning five broad data archetypes: tissue, cell culture, yeast, H&E, and bacteria. Because CellSAM can segment different cellular targets across image modalities, it can be immediately applied across bioimaging analysis workflows without requiring task-specific adaptations, or requires only retraining with few labels for boosted performance if needed. For example, segmentations from CellSAM can be used to track cells and quantify fluorescent reporter activity in cell culture, or to track cells across video frames for accurate lineage construction and cell division quantification. CellSAM can also segment individual slices of a 3D image and aggregate them to construct 3D cell segmentations. Additionally, CellSAM can be integrated with spatial transcriptomics pipelines like Polaris, where accurate cell segmentation allows assigning detected mRNA transcript spots to specific cells based on which cell boundary they fall within. A user interface for CellSAM is available at https://cellsam.deepcell.org/.
Read Paper | Nature Methods
4. Protein Embedding Compression. Discretizing protein embeddings would enable autoregressive generation using LLM infrastructure, potentially enabling unified models that both understand and generate protein content. However, the dimensionality of current protein embeddings makes direct discretization computationally intractable.
Lu et al. developed CHEAP, a method that substantially compresses the latent space of ESMFold (a protein structure prediction model built on the ESM2 protein language model) into continuous compressed embeddings, which can then be discretized into token representations. Since ESMFold maps from sequence to structure, its intermediate latent space can be viewed as a representation of the joint distribution of protein sequence and structure. The study systematically investigates how much compression can be applied and found that the latent space can be reduced by 128× in channel dimension and 8× in length while retaining structural information at <2Å accuracy, revealing that structural information is harder to compress than sequence information. For discrete tokenization, they show that FSQ (Finite Scalar Quantization) outperforms the commonly used VQ-VAE approach, especially for larger codebooks. Unlike concurrent structure tokenization methods that require 3D structural input, CHEAP can derive structure-aware tokens from protein sequence alone.
Read Paper | Patterns
-- Emma Chen, Pranav Rajpurkar & Eric Topol