Week 261
Virtual Multiplex Immunofluorescence, Ophthalmology Foundation Model, LLMs for Immunology, Radiology Foundation Model, Policy
In Week #261 of the Doctor Penguin newsletter, the following papers caught our attention:
1. Virtual Multiplex Immunofluorescence. The tumor immune microenvironment (TIME) critically impacts cancer progression and immunotherapy response. While routine H&E staining reveals cell morphology (shapes, structures, nuclei), it cannot identify which proteins are activated inside or on cell surfaces, which is information essential for understanding immune cell states and interactions. Multiplex immunofluorescence (mIF) can simultaneously visualize multiple protein markers while preserving spatial architecture, but its high cost and labor intensive workflow make large scale studies prohibitively expensive.
Valanarasu et al. developed GigaTIME, a multimodal AI framework that generates virtual mIF images from routine H&E pathology slides, enabling population-scale tumor immune microenvironment analysis. GigaTIME uses an encoder-decoder architecture that takes 256×256 pixel H&E patches as input and outputs 21 binary mIF channel predictions (activated/not activated for each protein marker). GigaTIME was trained on 40 million cells with paired H&E and mIF data, then applied to 14,256 patients from Providence Health (51 hospitals across seven states in the US), generating 299,376 virtual mIF slides spanning 24 cancer types and 306 subtypes. This virtual population uncovered 1,234 statistically significant associations linking proteins, biomarkers, staging, and survival. Such analyses were previously infeasible due to mIF data scarcity. The approach was independently validated on 10,200 TCGA patients, who had H&E slides and genomic/clinical data but no mIF images. Comparing association analyses between Providence and TCGA revealed strong concordance (Spearman correlation 0.88), though the Providence dataset identified 33% more significant associations, highlighting the value of large-scale real-world data.
Read paper | Cell
2. Ophthalmology Foundation Model. Optometrists and general practitioners are often the initial contacts for patients with vision issues, yet their referral accuracy varies widely from 48.2% to 88.9% for optometrists and 33% to 67.2% for general practitioners. Can AI reduce such variation and compensate for the scarcity of skilled practitioners in eye screenings and rural areas?
Soh et al. developed Meta-EyeFM, a system that combines a large language model with eight specialized vision foundation models to enable conversational diagnostics for primary eye care. Meta-EyeFM uses an "embedding-as-router" paradigm that dynamically directs user queries and fundus photographs to the most appropriate vision model trained to detect major eye diseases; differentiate ocular disease severity; predict systemic diseases (for example, 34.6% of people with diabetes are affected by diabetic retinopathy to some degree, but many are unaware due to its insidious nature); and identify specific ocular signs. The system outperforms both Gemini-1.5-flash and GPT-4o by 11-43% in disease detection and generally surpasses junior ophthalmologists and optometrists in clinical benchmarking, particularly excelling at glaucoma detection.
Read Paper | Cell Reports Medicine
3. LLMs for Immunology. Systems vaccinology seeks to understand vaccine-induced immunity by integrating large-scale molecular data with cellular and functional immune readouts. This biological complexity, coupled with strong context dependence across age, environment, and pathogen exposure, raises a critical question: can large language models (LLMs) engage in creative scientific reasoning within such a mechanistically complex field?
Rodriguez-Coffinet et al. evaluated whether LLMs can engage in genuine scientific discovery within the complex field of systems vaccinology using "The Creation Game," a framework inspired by the Turing test designed to assess AI's capacity for human-like scientific creativity. They tested five LLMs (ChatGPT-4o, ChatGPT-4.5, Microsoft Copilot, SciSpace, and LLaMA) using three mechanistically distinct case studies involving GCN2, SREBP, and TLR5 pathways in vaccine responses. The Creation Game goes beyond testing factual accuracy by evaluating whether AI can replicate the full arc of scientific reasoning: surveying literature, formulating hypotheses, designing validation experiments, and inferring broader biological principles. While the models demonstrated strong capabilities in factual recall, pattern recognition, and structured hypothesis formulation, they largely reorganized established findings rather than generating truly novel insights worthy of empirical testing. The authors emphasize that combining LLMs with algorithmic reasoning frameworks could enhance their ability to formulate biologically grounded insights, and they advocate for a partnership between human expertise and AI analysis as "hybrid intelligence" that maintains rigorous validation, ethical governance, and human oversight, ensuring AI serves as a catalyst for scientific inquiry rather than a constraint on creativity.
Read Paper | Science Immunology
4. Radiology Foundation Model. Medical volumes are immense in both spatial scale and bit-depth, making it challenging to model volumetric medical imaging. Consequently, leading foundation models from Google (MedGemma), Microsoft (MedImageInsight), and Alibaba (Lingshu) process CT exams as independent 2D slices, ignoring volumetric structure and losing essential contextual information.
Agrawal et al. developed Pillar-0, a radiology foundation model that processes full-resolution 3D CT and MRI scans. Trained on over 155,000 scans, including abdomen-pelvis, chest, and head CTs plus breast MRIs, it outperforms MedGemma, MedImageInsight, Lingshu, and Merlin (Stanford) by 7.8-15.8 AUROC points. Unlike existing methods that process CT scans as 2D slices with downsampled 8-bit images, Pillar-0 preserves native 3D volumetric structure and full 12-16 bit intensity information through the Atlas vision transformer architecture and multi-window tokenization. The Atlas architecture uses efficient multi-scale attention that enables processing of full 3D volumes (running 175× faster than standard vision transformers) while multi-windowing preserves clinically relevant intensity information across multiple tissue types. Additionally, the model achieves 20-40× better data efficiency than competing approaches, reaching 95% accuracy on brain hemorrhage detection using just 1/20th the training data. The study also developed RATE, a unified framework designed to evaluate any vision model on full-fidelity medical volumes using authentic clinical tasks derived from real-world radiology practice, by applying large language models to extract answers from unstructured radiology reports for a set of clinically grounded queries. The team has released all model weights, training code, and evaluation frameworks as open-source resources.
Read Paper | arXiv
5. Policy. How do AI health tools actually get paid for in the U.S. healthcare system?
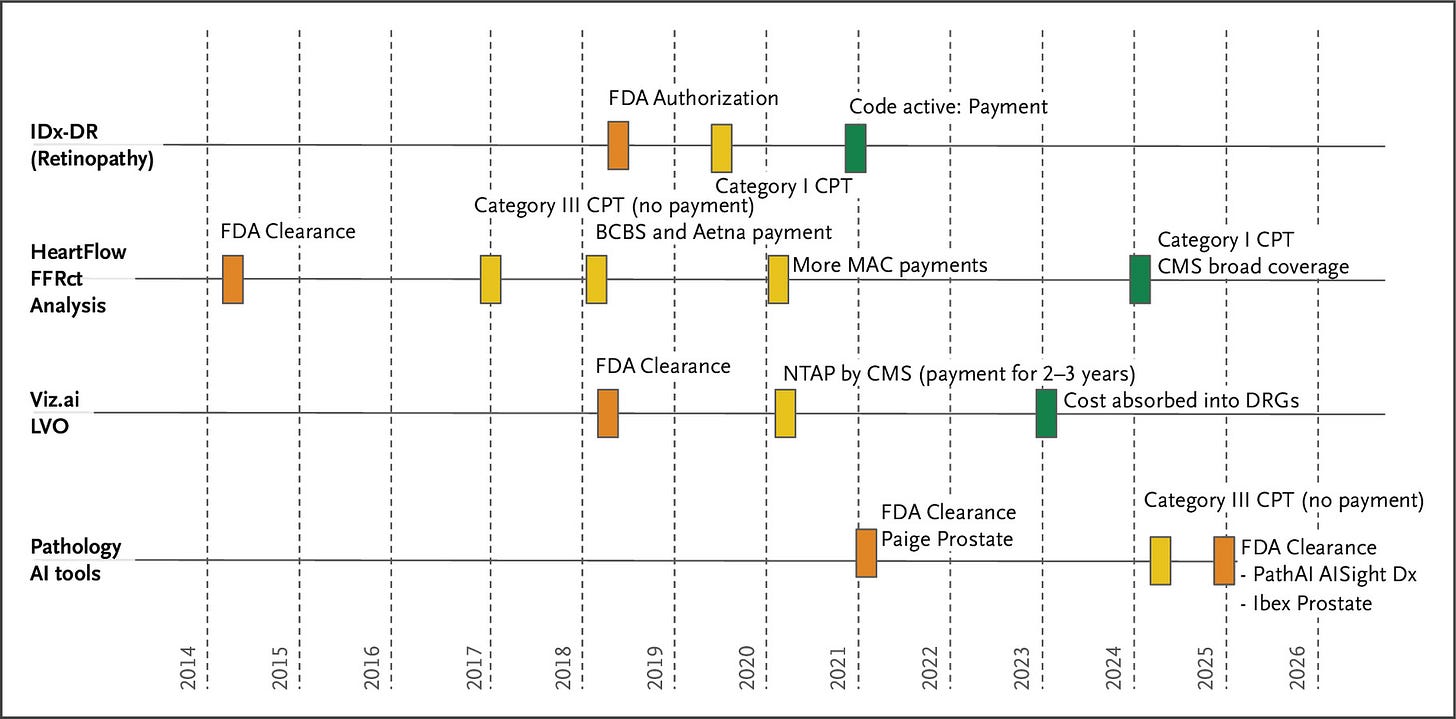
In this article, Razavian et al. examine why healthcare remains largely untransformed by AI despite rapid advances across other sectors, identifying payment system barriers as the primary obstacle to widespread adoption. In the United States, payment mechanisms fall into three categories: inpatient bundled payments, outpatient bundled payments, and fee-for-service payments governed by CPT codes. Most AI products pursue the CPT pathway through a lengthy sequence: obtain FDA clearance, partner with a medical specialty society to petition the AMA's CPT Editorial Panel, receive either a Category III code (temporary, no payment, tracks utilization for up to 5 years) or Category I code (permanent, eligible for payment, requires "widespread use"), undergo pricing assignment by the AMA's RUC committee, secure CMS coverage determinations, and finally await independent coverage decisions from commercial insurers. Recognizing this process is too slow, CMS created temporary pathways, NTAP for inpatient and TPT for outpatient settings, providing supplemental payment for up to 3 years while data accumulates. Through case studies, the authors show that even FDA-cleared AI tools with demonstrated clinical efficacy face massive adoption barriers, with payment approval typically taking 5-10 years after FDA clearance due to outdated frameworks and misaligned incentives across hospitals, insurers, physicians, and AI companies. The authors propose five policy recommendations: streamline AI coding by categorizing services by function rather than treating each tool uniquely; eliminate the "widespread use" catch-22 by allowing provisional payment for FDA-cleared tools with clinical efficacy; address API integration costs by requiring EHR vendors to provide free data access; create flexible pricing that adapts to AI's rapidly declining costs; and foster competitive markets by bundling similar AI tools and sharing performance data.
Read Paper | NEJM AI
The Timeline and Major Reimbursement Events in a Few Health AI Domains with U.S. Food and Drug Administration–Approved Products. Source: Razavian et al., NEJM AI 2025.
-- Emma Chen, Pranav Rajpurkar & Eric Topol