We are moving our newsletter to Substack for a better experience!
In Week #217 of the Doctor Penguin newsletter, the following papers caught our attention:
1. Foundation Model. Foundation model for single-cell multi-omics.
Drawing parallels between language and cellular biology where texts comprise words and cells are defined by genes, Cui et al. developed a transformer-based foundation model for single-cell biology called scGPT that simultaneously learns cell and gene representations. scGPT was trained on single-cell sequencing data of 33 million normal human cells from 51 organs or tissues. During training, the model gradually learns to generate gene expression of cells based on cell states or gene expression cues. scGPT achieves state-of-the-art performance on downstream tasks including cell type annotation, genetic perturbation prediction, batch correction, and multi-omic integration after fine-tuning. Additionally, comparisons of gene embeddings and attention weights between fine-tuned and raw pre-trained models uncover valuable biological insights into gene-gene interactions specific to cell types and perturbation states. Moreover, this study observes a scaling effect: larger pretraining data sizes yield superior pretrained embeddings and further lead to improved performance on downstream tasks.
Read paper | Nature Methods
2. Large Language Model. How well can large language models (LLMs) analyze and summarize clinical text?
Van Veen et al. evaluated methods for adapting LLMs to clinical summarization tasks. The adaptation methods included (1) in-context learning, which adapts the model by including examples within the prompt, and (2) quantized low-rank adaptation, which adapts by fine-tuning a subset of model weights on examples. They applied these methods to 8 LLMs across 4 distinct clinical summarization tasks (radiology reports, patient questions, progress notes, and doctor-patient dialogue). Using quantitative NLP metrics, they identified the best model and adaptation method for each task. Next, they conducted a clinical reader study with 10 physicians comparing the best LLM summaries to expert summaries on completeness, correctness and conciseness. The study found LLM summaries were often preferred over expert summaries due to higher scores on those key attributes. Finally, they performed a safety analysis of examples, potential medical harm and fabricated information to understand the challenges faced by both models and medical experts. Overall, this study suggests incorporating LLM-generated candidate summaries could reduce clinician documentation burden, potentially leading to decreased clinician strain and improved patient care.
Read Paper | Nature Medicine
3. Large Language Model. Large language models (LLMs) in ophthalmology.
Huang et al. conducted a single-center, cross-sectional study with 15 ophthalmologists (12 attending physicians and 3 senior trainees) to compare the diagnostic accuracy and comprehensiveness of responses from GPT-4 versus fellowship-trained glaucoma and retina specialists. Randomly selected glaucoma and retina questions and de-identified patient cases were used to compare answers to clinical questions and case management plans generated by GPT-4 and the specialists. GPT-4 demonstrated proficiency comparable to, if not better than, the specialists. The chatbot's strong performance may stem from the refined prompting techniques used, specifically instructing the model to respond as a clinician in an ophthalmology note format. These findings highlight the potential for LLMs to serve as valuable diagnostic aids in ophthalmology, especially in highly specialized subspecialties like glaucoma and retina surgery.
Read Paper | JAMA Ophthalmology
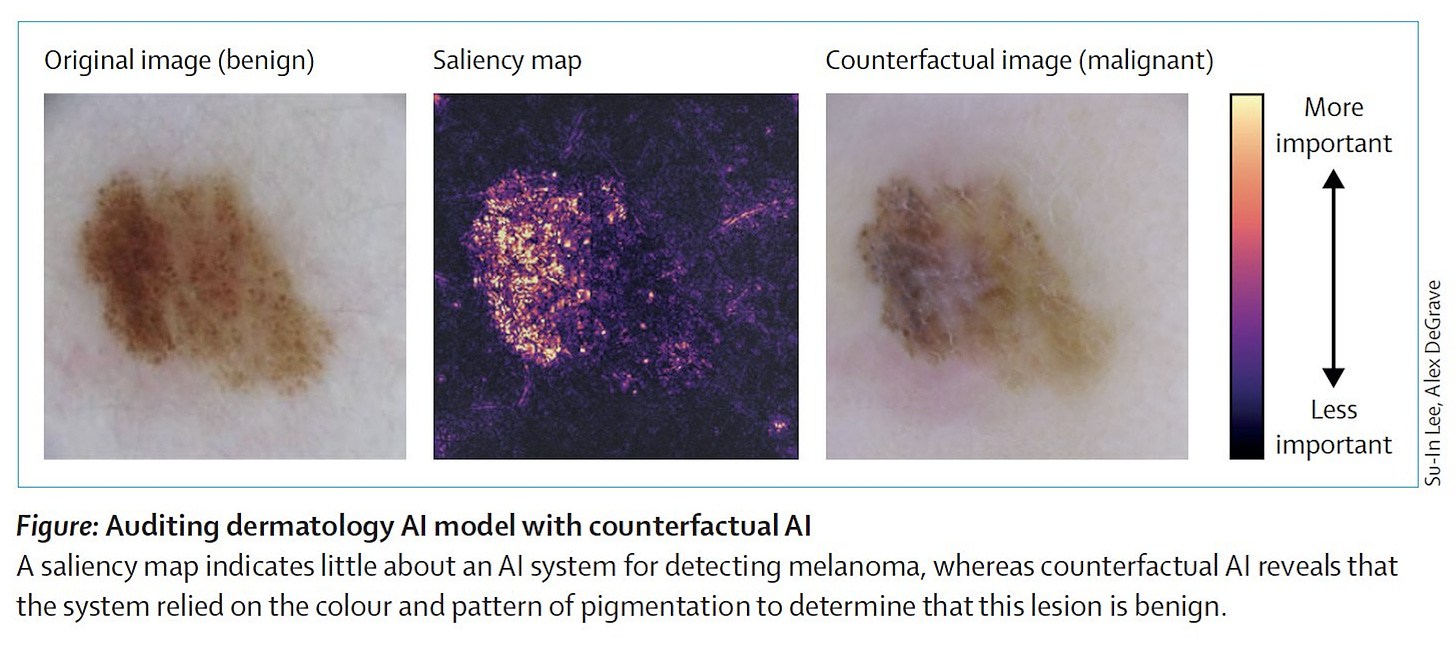
4. Counterfactual Al. “Generative Al informs the development of explainable Al; explainable Al aids in understanding generative Al models.”
In this Perspective, Lee and Topol highlight a new data-centric approach that uses generative AI to generate counterfactual data to explore the "what if" scenarios in research, which they term "counterfactual AI." This method can be used to study how patient data might change under specific conditions, fill data gaps for rare diseases or underrepresented groups, and aid drug discovery and development with the generated data. More importantly, counterfactual AI expands the scope of explainable AI by providing counterfactual data that elicit specific outcome predictions from complex AI classifiers, allowing humans to gain more comprehensive insights into the reasoning processes of these classifiers.
Read Paper | The Lancet
-- Emma Chen, Pranav Rajpurkar & Eric Topol